On October 20, 2015, we published a blog post about the novel functionalities in ARPHA that allows streamlined import of specimen or occurrence records into taxonomic manuscripts.
Recently, this process was reflected in the “Tips and Tricks” section of the ARPHA authoring tool. Here, we’ll list the individual workflows:
- Upload occurrence or specimen data from a Darwin-Core-compliant Excel spreadsheet
- Upload occurrence or specimen data manually
- Upload occurrence or specimen data from the GBIF repository
- Upload occurrence or specimen data from the BOLD Systems repository
- Upload occurrence or specimen data from the iDigBio repository
- Upload occurrence or specimen data from the PlutoF repository
Based on our earlier post, we will now go through our latest updates and highlight the new features that have been added since then.
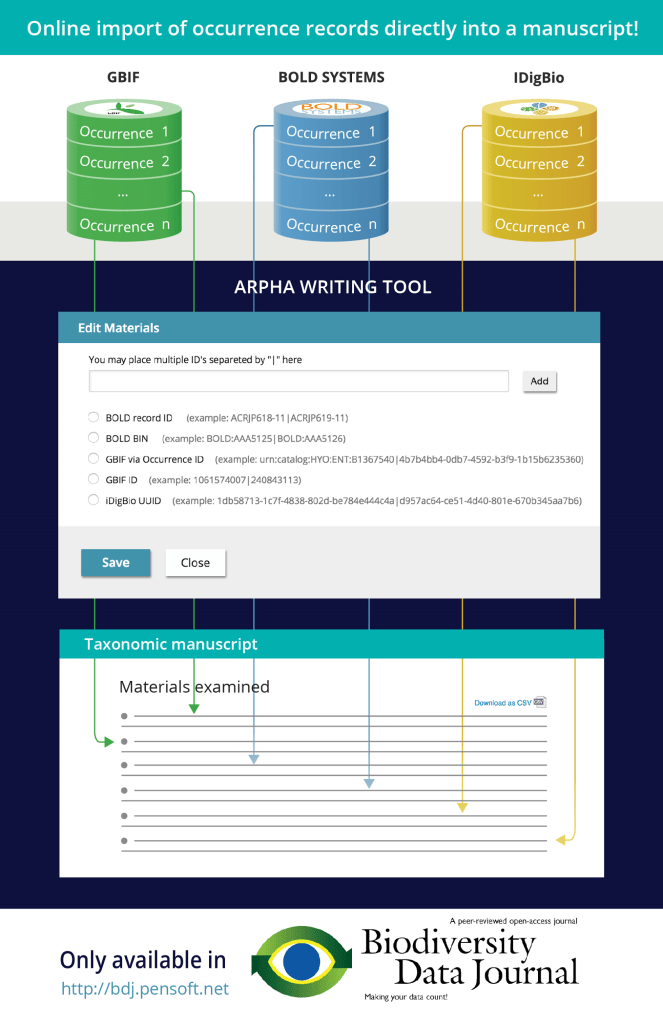
Repositories and data indexing platforms, such as GBIF, BOLD systems, iDigBio, or PlutoF, hold, among other types of data, specimen or occurrence records. It is now possible to directly import specimen or occurrence records into ARPHA taxonomic manuscripts from these platforms [see Fig. 1]. We’ll refer to specimen or occurrence records as simply occurrence records for the rest of this post.

The functionality shows up when one creates a new “Taxon treatment” in a taxonomic manuscript in the ARPHA Writing Tool. To import records, the author needs to:
- Locate an occurrence record or records in one of the supported data portals;
- Note the ID(s) of the records that ought to be imported into the manuscript (see Tips and Tricks for screenshots);
- Enter the ID(s) of the occurrence record(s) in a form that is to be seen in the “Materials” section of the species treatment;
- Select a particular database from a list, and then simply clicks ‘Add’ to import the occurrence directly into the manuscript.
In the case of BOLD Systems, the author may also select a given Barcode Identification Number (BIN; for a treatment of BIN’s read below), which then pulls all occurrences in the corresponding BIN.
We will illustrate this workflow by creating a fictitious treatment of the red moss, Sphagnum capillifolium, in a test manuscript. We have started a taxonomic manuscript in ARPHA and know that the occurrence records belonging to S. capillifolium can be found on iDigBio. What we need to do is to locate the ID of the occurrence record in the iDigBio webpage. In the case of iDigBio, the ARPHA system supports import via a Universally Unique Identifier (UUID). We have already created a treatment for S. capillifolium and clicked on the pencil to edit materials [Fig. 2].


Not having to retype or copy/paste species occurrence records, the authors save a lot of efforts. Moreover, they automatically import them in a structured Darwin Core format, which can easily be downloaded from the article text into structured data by anyone who needs the data for reuse.
Another important aspect of the workflow is that it will serve as a platform for peer-review, publication and curation of raw data, that is of unpublished individual data records coming from collections or observations stored at GBIF, BOLD, iDigBio and PlutoF. Taxonomists are used to publish only records of specimens they or their co-authors have personally studied. In a sense, the workflow will serve as a “cleaning filter” for portions of data that are passed through the publishing process. Thereafter, the published records can be used to curate raw data at collections, e.g. put correct identifications, assign newly described species names to specimens belonging to the respective BIN and so on.
Additional Information:
The work has been partially supported by the EC-FP7 EU BON project (ENV 308454, Building the European Biodiversity Observation Network) and the ITN Horizon 2020 project BIG4 (Biosystematics, informatics and genomics of the big 4 insect groups: training tomorrow’s researchers and entrepreneurs), under Marie Sklodovska-Curie grant agreement No. 642241.